Authors/Affiliations:
The concept of Open Bibliography in science, technology and medicine (STM) is introduced as a combination of Open Source tools, Open specifications and Open bibliographic data. An Openly searchable and navigable network of bibliographic information and associated knowledge representations, a Bibliographic Knowledge Network, across all branches of Science, Technology and Medicine, has been designed and initiated. For this large scale endeavour, the engagement and cooperation of the multiple stakeholders in STM publishing - authors, librarians, publishers and administrators - is sought.
BibJSON, a simple structured text data format (informed by BibTex, Dublin Core, PRISM and JSON) suitable for both serialisation and storage of large quantities of bibliographic data is presented. BibJSON, and companion bibliographic software systems BibServer and OpenBiblio promote the quantity and quality of Openly available bibliographic data, and encourage the development of improved algorithms and services for processing the wealth of information and knowledge embedded in bibliographic data across all fields of scholarship.
Major providers of bibliographic information have joined in promoting the concept of Open Bibliography and in working together to create prototype nodes for the Bibliographic Knowledge Network. These contributions include large-scale content from PubMed and ArXiv, data available from Open Access publishers, and bibliographic collections generated by the members of the project. The concept of a distributed bibliography (BibSoup) is explored.
This paper "eats its own dog food" by using the technologies described in the text. All bibliographic entry references and bibliographic entries were managed in BibJSON then included in the HTML document following the Scholarly HTML convention. The document itself is formally consistent with these specifications and can be read as a normal HTML document. It would alternatively be possible to embed bibliographic records in the document directly from BibJSON via JavaScript. The "flat HTML" should be taken as the definitive version, and can be re-purposed into other formats (PDF, DOCX, ODT).
We introduce the concept of Open Bibliography as a combination of Open Source tools, Open specifications and Open bibliographic data. Our Open Bibliography project is an umbrella of several other initiatives, most prominently the Open Knowledge Foundation’s Bibliographic Working Group [1], the JISC-funded JISC-OpenBib project at the University of Cambridge [2], and the NSF-funded Bibliographic Knowledge Network project [3]. These projects have all addressed the totality of Open bibliographic resources including design of systems, implementation of software, licenses for use and re-use, and the collection and hosting of substantial bibliographic datasets. In this article we shall concentrate on bibliographic data for articles in the Science, Technology and Medicine (STM) fields, but we introduce the reader to the wider elements of bibliography before the main results. We stress that the tools and formats exemplified here have a particularly simple modular form in STM article publishing; however, these tools and formats are designed to be both flexible and extensible, and are also capable of managing library and personal collections, monographs, multiple versions etc.
The atoms in the bibliographic universe are the records traditionally held as 3 X 5 cards in a library card catalogue, and more recently represented in structured text languages such as BibTeX, RIS, XML, JSON [4] [5] [6] [7]. This data has been commoditised, and is subject to a large scale cycle of use (publication / collection / abstracting / indexing / searching / citation) which involves all participants in scientific publication: authors, librarians, publishers and administrators.
The bibliographic data cycle is reminiscent of the water cycle, whereby water stored in the atmosphere rains down onto the land, where it may be collected in containers on various scales (drops, thimbles, buckets, ponds, and lakes), from which it flows down rivers and pipes to the ocean, thence back to the atmosphere by evaporation to close the cycle.
Similar features of the bibliographic data cycle are the variety of containers of different sizes that can be used to enclose it (individual records, publication lists, departmental collections, subject-specific repositories, and the databases behind large scale indexing services), the use of the internet to pipe data from one place to another, the way bibliographic items are recycled in the process of bibliographic citation, and the way bibliographic data is packaged and sold to universities. But there is a key difference which mitigates the prevailing business model of bibliographic data as a commodity controlled by bibliographic service providers and sold to universities at a substantial profit; unlike volumes of a liquid, bibliographic data is not subject to any conservation law. Rather, bibliographic data is subject to a process of continual creation and replication. The elements of bibliographic data are facts, which in most jurisdictions cannot be copyrighted. There are few technical and legal obstacles to widespread replication of bibliographic records on a massive scale - the main limitations of such activity are social: whether individuals and organisations are adequately motivated to create and maintain open bibliographic resources. But the dynamics of creation and replication of bibliographic records have been irreversibly changed by recent technological and social developments, most notably the emergence of:
Bibliographic data has long been understood to contain important information about the the influence and impact of various authors and journals on scientific disciplines [18] [19]. However, now, instead of a relatively small number of privileged data owners being able to manage and control large bibliographic data stores [20], an individual researcher could browse millions of records, view collaboration graphs, submit complex queries, make selections and analyses of data – all on their laptop while commuting to work, provided they have adequate tools to do so.
The tools for such easy processing are not yet adequately developed, so our aim is to provide Open tools and services to make the wealth of bibliographic data available to the widest possible audience, and to promote increased understanding of science and technology, especially in interdisciplinary areas.
Traditionally, bibliography has been regarded as the study of library holdings and catalogues, and, more recently, catalogues of material published by formal publishers, repositories and other collections. We wish therefore to explain the importance of bibliography to scientists, and to argue the merits of Open Bibliography, by which we mean systematic efforts to create and maintain stores of Openly accessible [21], machine-readable bibliographic data.
The unit of bibliography is the bibliographic record, which consists of the information necessary to locate and/or identify a scholarly publication (and, increasingly, other resources besides textual material, such as authors, images and scientific datasets). The term "bibliography" is also often used to represent a personal collection of bibliographic records (and in some cases is synonymous with citation lists). We refer to such a collection here as a "bibliographic dataset".
We use the term "citation" to mean a reference to a bibliographic record within the body of a document. A citation may also be called simply a "reference". We will not say much more about citations, except to point out that an improved approach to bibliography should also be of value to citation management and analysis.
Despite the importance of bibliography, including the widespread sale of bibliographic records, there is no single syntax or agreed semantics for the publication and exchange of STM bibliographic records; scientists use whatever representation is provided by their tools, the most Open and common being BibTeX from the LaTeX [22] authoring system; the publishing community commonly uses PRISM, although some publishers have their own representation of their bibliographic data, which often consists of a mixture of Dublin Core [23], PRISM, BibTeX and their own markup approach.
By "Open Bibliography", we do not imply that there must be a single central authoritative resource. We expect that the first stage, at least, will be the identification of bibliographic collections which are Open and where the collectors can offer them with the appropriate common technology. The aggregation of such distributed collections, with some cooperation about formats and interfaces, is what we mean by the Bibliographic Knowledge Network.
We are aware of other groups with explicit commitments to use of open licenses such as K4All [24] and also Wikipedia [25] which has a large implicit bibliography. There are also a substantial number of large bibliographic repositories which are operationally fairly Open, even if the data is not explicitly declared to be so, or available only through an API rather than in bulk, such as arXiv, RePEc, BibSonomy, PhilPapers, DBLP, CiteULike, Connotea, Zotero, Mendeley etc. [26] [27] [28] [29] [30] [31] [32] [33] [34]. We have worked closely with two major Open Access publishers (International Union of Crystallography (IUCr) and BioMed Central (BMC)) [35] [36], and have collaborated with PubMed [37], with Thomas Krichel’s AuthorClaim [38] and 3Lib [39] projects, and with the Sciplore team [40].
Because this article is limited to exploring bibliography for STM, we have taken a pragmatically simple approach. Systems such as FRBR [41] and BIBO [42] make provision for complex aspects of bibliography such as multiple manifestations and representations of works and multiple versions. While these are relevant to STM bibliography in some areas [43], for the most part we do not need the complexity of these and other RDF [44] approaches, although our tools and software should be capable of leveraging them if necessary.
We report a number of prototypes in both tools and collections, and also propose that STM bibliography can be adequately represented for most immediate purposes using BibJSON [45]. Due to the intense interest in Open Bibliography, we are now very actively working on future versions of BibJSON, but the examples given in this article are fully supported by current software. In the spirit of the "perpetual beta" approach on the web, we intend to release early and often in public view so that a broad community becomes intimately involved in the design of specifications. As a first example, the references in this document are stored in a prototype of BibJSON and can be rendered into the content via JavaScript.
We present in this section a list of reasons and use cases which motivate our commitment to Open Bibliography.
Bibliography, and bibliographic data, is sometimes regarded as referring to everything that is not part of the "full text" and "images" in an article. This can be problematic because some people and organisations regard material such as abstracts, annotations and citation lists as "copyrightable" and therefore not by default Open. In this article, we do not debate the ethics and legality of asserting ownership over certain types of bibliographic data, and our understanding of the agreed law and practice is that what we define as "core bibliographic data" below can be made Open by default.
By "core bibliographic data" we mean that data which is necessary to identify and / or discover a publication. It is generally held that such bibliographic data is NOT copyrightable and this has been confirmed by the Association of STM Publishers in a public reply to one of the authors [53].
It is difficult to get authoritative statements as to whether other fields are Open by default. But we would expect, for example, that the format of the work and the rights associated with it were by default Open, while the abstract and images were not. Traditionally collections of STM bibliographic data have been expensive to produce and most of these are therefore currently available only under licenses that restrict re-use. Because it is now technically possible to create large amounts of Open Bibliographic data, this opens the possibility of collections created from the start as Open and distributed for community re-use.
The following "core bibliographic data", as described by the Open Bibliography Principles [54], will be the subject of this article:
A number of ways of creating Open Bibliographic data may be identified:
Using these and other mechanisms, we believe that is it cost effective to create and maintain an Open bibliographic network of information about STM publications. These need not necessarily be electronic publications but the stress of this article will be on the collection of bibliographic data that refers to electronic journals, web pages, technical reports, theses, and documents available on the web, meaning the data that is required to locate and identify a document on the web, whether or not the full text of that document may is openly available. As an example, it is possible to extract bibliographic data for all the publications in the BMC collection of journals. The web has been crawled for many years and the technology for doing this is standard. It is polite, but not legally required, to agree large-scale crawling with a publisher or to create web-server-friendly robots which do not impose undue stress.
Most scientists require a single bibliographic record per publication. In other words, most scientists do not distinguish between a print version, an electronic version or a manuscript on an author’s web page or in their institutional repository. Scientists have the implicit model of a single platonic bibliographic record for an article. Our approach is based on this and while there may be occasional complexities that cannot be represented, we believe it is powerful enough to create a useful sustainable Open STM bibliography.
The vocabulary terms used by publishers and other bibliography creators, often drawn from Dublin Core, PRISM, Medline or home grown element sets, are fairly, but not completely, interchangeable. For example. dc:creator might be used for authors in one source and editors or publishers in other sources, but usage is normally consistent within a given source. As a first step we propose to honour the terms used by the collectors rather than attempt to align and normalise them algorithmically. We are exploring whether there is a pragmatic "flattening" of the main concepts and whether it is possible to manage "most" STM bibliographic records with a small number of central terms; most STM articles in journals can be described with a very small subset of these vocabularies:
| Types of entities | ||
|---|---|---|
| Name | Element Set(s) | Description |
| Agent | FOAF, dcterms | A resource that acts or has the power to act. |
| Person | FOAF | A person |
| Organisation | FOAF | An organisation |
| Document, Bibliographic Resource | FOAF, BIBO, dcterms | A document of some sort |
| Article | BIBO | An article, typically in a Journal |
| Issue | BIBO, BibTeX | A journal issue or volume (expressed as a property in BibTeX, linked with dcterms:isPartOf in BIBO) |
| Journal | BIBO, BibTeX | A journal (expressed as a property in BibTeX, linked with dcterms:isPartOf in BIBO) |
| Properties or predicates | ||
|---|---|---|
| Name | Element Set(s) | Description |
| author, creator, contributor,editor | BibTeX, dcterms | Person or organisation creatively responsible for some document |
| identifier | BibTeX, BIBO, dcterms | Identifier of an entity such as an article or journal (including refinements such as ISSN, DOI, ISBN, etc. which are common BibTeX extensions) |
| institution | BibTeX | The institution involved in publishing |
| journal | BibTeX | A journal (see Journal in classes above) |
| month, year, published | BibTeX, dcterms | The date of publication |
| name, label | FOAF, RDFS, SKOS [57] | A name or label for a thing such as a person or organisation. |
| pages, extent | BibTeX, BIBO, dcterms | Page numbers |
| publisher | BibTeX, dcterms | A publisher |
| title | BibTeX, dcterms | The title of the work |
| volume | BibTeX | The volume of a journal (see Issue in classes above) |
We could equally well have included the relevant fields from MARC21 and more [58] [59] in the above table. What these representations, MARC21, BibTeX and BIBO+dcterms, have in common is a flat representation of a bibliographic record. Contrast with the WEMI model of FRBR where a single bibliographic record for an article would be separated out into three or four related entities, Work, Expression, Manifestation at least, according to complicated cataloguing rules, without even considering the added dimensions of journals, authors and publishers. This flat representation is a core feature of our conceptual model.
Identifiers are critically important. They are necessary (but obviously not sufficient) to enable tasks like de-duplication - in order to identify duplicates, we need to be able to identify the things that are duplicated. They also make it possible to refer to entities outside of the current dataset; one might refer to the author of an article by their Wikipedia page, for example. This is not necessary, but it opens up many interesting possibilities for interlinking and correlating amongst datasets. Using a URI as an identifier where feasible is therefore a desirable feature [60].
Where a single, sustainable resource manages bibliographic data, it makes sense for it to generate its own unique identifiers, even if there is already a well-defined identifier system for some of the information. Thus, in working with the British Library [61] on the British National Bibliography [62], we have created a set of identifiers for their records. However, where collections come from several sources it is very difficult to create a global unique identifier system without a curating organisation. We therefore expect that each collection will create its own identifiers. We expect that different collections will contain bibliographic data for the same object and here we will create a mapping between the collections rather than trying to create a single global index.
We have also worked with the following datasets (see also Section 6) and found that the records can be well represented by the concepts above.
With this conceptual model in hand, we can turn our attention to exchanging information between systems that have similar or at least compatible models. For pragmatic reasons we propose to use JSON to exchange this information. JSON is widely implemented, simple to parse and easy to create either with a computer program or by hand in a text editor. A JSON-based format which uses dictionaries or associative arrays is also extensible since adding a new key to such a dictionary should not break any existing implementations which may not understand the meaning of the new key.
By using a JSON-based format designed for representation of bibliographic data - meaning data about documents of various kinds, and about the people, organizations and subjects connected to those documents - we can include guidance for creating records, linkages to existing ontologies, vocabularies and schema, and schema definitions, covering a wide range of bibliographic needs and drawing from a number of bibliographic metadata sources (BIBO, BibTeX, DC). If desired, a creator of a bibliographic dataset may add more information (e.g. language, format, editors, etc.). A consumer of this dataset may or may not read and understand this.
This paper is strongly informed by the work done in the JISC-OpenBib project, a collaboration between the University of Cambridge, the OKF, the British Library, Cambridge University Library and the IUCr. With the help of these partners, the high-level goal was to take exemplary bibliographic datasets and show that the principles of Open Bibliography, coupled with the formalisation and tools reported here, would be of great value to the scientific and informatics communities. We report a number of successful prototypes in this section where we have been able to acquire or collect an Open dataset, transform it to BibJSON or equivalent and re-purpose it, often with an interactive tool. The emphasis in this paper is on STM bibliographic resources but for completeness we also report on other collections.
The OKF has developed a bibliographic management system (Bibliographica) which was functional at the start of this project. Although general, it has been primarily aimed at non-STM resources such as library collections and personal bibliographies. During the project, the British Library released under a CC0 licence the British National Bibliography (a collection of about 3 million records for monographs created in their role as deposit library). These have been converted to as queryable RDF using our Open Source OpenBiblio software [65] [66]. This provides an example of using the software to make bibliographic metadata available as RDF where required; this and other instantiations of OpenBiblio then act as resources for building bibliographic collections.
The largest collection of STM bibliographic data is provided by the NLM from the National Institutes of Health (NIH) of the USA. This is provided freely, and the records refer to both Open and non-Open publications. The Open publications are referred to as the "Open Access subset (OAS)" (the terminology is complex and our project has explained it [67] ). The OAS (ca. 250,000 records) contains full text and full reference lists (citations), and is of a very tractable size for carrying out prototypic work on bibliography and citations. The full Medline collection has about 20 million articles and in collaboration with the NLM we have obtained these records and converted them to RDF using a straightforward BIBO+dcterms representation. For the full record set we have been careful to include only those components of bibliography which are agreed to be Open (i.e. we have omitted abstracts and editorial annotation). Nevertheless, this collection is a major new resource in Open Scholarship.
We have converted both subsets to RDF, and found that while the Open subset is tractable with a wide range of common tools, the full records have problems of scale. It produced over 1 billion RDF statements; the resources required for querying this in an RDF store are beyond current scope, however a sample record is appended, and further information along with full content is Openly available [68] [69]. For the full subset we are now using a BibJSON-like approach and storing the records in a NoSQL database (CouchDB). This gives good performance for the sorts of queries that most people will initially wish to make.
Despite not having the abstracts or full text Openly, the Medline bibliographic dataset has still enormous value, particularly when used with new ways of navigation and display.
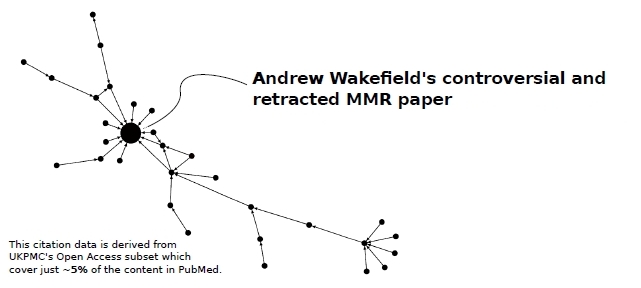
Although citations (bibliographic entry reference lists) are outside the scope of Open Bibliography, the OAS provides an opportunity to work on citations. This is less easy because the reporting of citations is poorly formalised (a major motivation for Open Bibliography and BibJSON), and they contain a large number of errors, including non-existent bibliographic objects. However, the potential is large and we display an analysis of citations related to a retracted paper [70].
Figure 1: A citation map of papers recursively referencing Wakefield's paper on the adverse effects of MMR vaccination. A full analysis requires not just the act of citation but the sentiment, and initial inspection shows that the immediate papers had a negative sentiment i.e. were critical of the paper. Wakefield's paper was eventually withdrawn but the other papers in the map still exist. It should be noted that recursive citation can often build a false sense of value for a distantly-cited object.
Traditionally, bibliographic records have been seen as a management tool for physical and electronic collections, whether institutional or personal. In bulk, however, they are much richer than that because they can be linked, without violation of rights, to a variety of other information. The primary objective axes are:
With these components, it is possible to create a living map of scholarship, and we show two examples carried out with our bibliographic sets.
Figure 2: This is a geo-temporal bibliographic map for crystallography. The IUCr's Open Access articles are an excellent resource as their bibliography is well-defined and the authors and affiliations well-identified. The records are plotted here on an interactive map where a slider determines the current timeslice and plots each week's publications on a map of the world. Each publication is linked back to the original article. (The full interactive resource is available at http://benosteen.com/timemap/index.)
Figure 3: This is a geo-temporal bibliography from the full Medline dataset. Bibliographic records have been extracted by year and geo-spatial co-ordinates located on a grid. The frequency of publications in each grid square is represented by vertical bars. (Note: Only a proportion of the entries in the full dataset have been used and readers should not draw serious conclusions from this prototype). (A demonstration screencast is available at http://vimeo.com/benosteen/medline; the full interactive resource is accessible with Firefox 4 or Google Chrome, at http://benosteen.com/globe.)
These visualisations show independent publications, but when the semantic facets on the data have been extracted it will be straightforward to aggregate by region, by date and to create linkages between locations.
Bibliographic data are particularly valuable for mashups (i.e. the combination of data components that share one or more common values or identifiers). Thus, for example, it should be possible to link Open Bibliography to bibliographic entry references in Wikipedia. More generally, Open Bibliography is available for any author or organisation who wishes a definitive identification of the bibliographic entry references in a document.
Our mashups demonstrate that when data is openly available, it enables serendipitous and relatively quick development of useful tools. For example, we created a Wikipedia bookmarklet, a personal collections tool on Bibliographica, and a relevant reading list generator for the Edinburgh International Science Festival [71] [72] [73].
In order to manage larger bibliographic datasets in a simpler format, we are now collaborating with Bibliographic Knowledge Network to develop the BibJSON format for representing bibliographic records. It is sufficient for most current purposes for basic STM articles, adequate also for all basic BibTeX types including monographs, and is extensible so it can easily support records for authors, journals, etc. The main virtues are:
We have created sample BibJSON conversions using our software (examples appended), and will continue to perform these conversions on the datasets now available. BibJSON can perform a similar function for communities wishing to share bibliographic data as GeoJSON [74] does for those sharing geospatial information.
The use of lightweight technology has allowed us to create a radically new approach towards the collection of Open bibliography. Conventional wisdom would suggest that all bibliographic records should be normalised and validated by a central authority. In BibJSON, we take the view that any Open bibliographic record (with its provenance) is potentially valuable, even though there may be duplicates referring to "the same bibliographic object". The question of determining whether two records relate to "the same object" is difficult and controversial and BibSoup deliberately avoids this. It consists of a number of collections of bibliography (initially in STM areas) united by a common syntax. It is left to humans and machines to develop annotations and equalities between the components of these collections. Thus, for example, "the same paper" will be reported in arXiv, DBLP and possibly even Medline.
The BibSoup approach encourages the contribution of Open bibliography without the overhead of de-duplication at contribution time. We expect that, as it grows, services will develop that help users and maintainers to manage the information. De-duplication into a central repository may be one solution (with the presumed platonic identity of STM bibliographic entries), but we also expect that software based on RDF will allow tools to manage alternative representations of bibliographic data, leaving the choice to the user as to what strategy they take. In short, current STM bibliography is a distributed mess. BibSoup takes this as a starting point and, where the political will and financial support is available, offers methods for tidying this up.
Via collaboration with the Scholarly HTML (ScHTML) [75] community we intend to follow conventions for embedding bibliographic metadata within HTML documents whilst also enabling collection of such embedded records into BibJSON and BibSoup, thus allowing embedded metadata whilst also providing additional functionality such as search. We are also working towards ensuring compatibility between ScHTML and Schema.org [76], affording greater relevance and usability of ScHTML data.
We are continuing development of BibServer [77] along with the BibJSON specification as a way for individuals - or departments or research groups - to easily manage, present, and search their own bibliographic collections. Collections can be stored in BibTex files, in JSON files or a JSON database such as CouchDB, or in an OpenBiblio instance, or managed directly by the software. The key to the architectural design is that it will be possible for other interested parties to develop their own plugins both for ingest and storage, allowing flexibility in implementation.
These ongoing efforts to develop OpenBiblio, BibJSON and BibServer, will enable us to support large scale Open Bibliographic data – the BibSoup. We hope to attract further collaborations from other groups which realise the importance of Open Source code, Open Data and Open Knowledge to the future of scholarship.
The following examples demonstrate conversions of typical bibliographic records into BibJSON. Although BibJSON is not a complete standard, the aim is to demonstrate the simplicity with which we can represent this data in a JSON object, using namespaces to extend keys as necessary. The default namespace for BibJSON keys is essentially BibTex plus a few keys required to support BibJSON, such as “namespaces”; anything beyond the scope of BibTex should be added by using a namespace.
IUCr raw bibliography:
<link rel="schema.DC" href="http://purl.org/dc/elements/1.1/ " /> <link rel="schema.DCTERMS" href="http://purl.org/dc/terms/ " /> <link rel="schema.prism" href="http://prismstandard.org/namespaces/1.2/basic/ " /> <meta name="DC.source" content="urn:issn:1600-5368" /> <meta name="DC.rights" content="http://creativecommons.org/licenses/ " /> <meta name="DC.creator" content="Zheng, L." /> <meta name="DC.creator" content="Hu, F." /> <meta name="DC.creator" content="Zeng, X.C." /> <meta name="DC.creator" content="Li, K.P." /> <meta name="DC.date" content="2011-04-01" /> <meta name="DC.identifier" content="doi:10.1107/by/2.0/uk S1600536811007148" /> <meta name="DC.publisher" content="International Union of Crystallography" /> <meta name="DC.link" content="http://scripts.iucr. org/cgi-bin/paper?cv5056 " /> <meta name="DC.language" content="en" /> <meta name="DC.description" content="The title compound, C11H14N2O5, was synthesized by condensation of (RS)-2-aminosuccinic acid dimethyl ester with 2-trichloroacetylpyrrole at room temperature. The amide group is twisted by 7.4 (1)degrees from the plane of the pyrrole ring. In the crystal, molecules are linked by intermolecular N-H...O hydrogen bonds into chains extending along the c axis." /> <meta name="DC.type" content="text" /> <meta name="DC.title" content="rac-Dimethyl 2-(1H-pyrrole-2-carboxamido)butanedioate" /> <meta name="DCTERMS.abstract" content="The title compound, C11H14N2O5, was synthesized by condensation of (RS)-2-aminosuccinic acid dimethyl ester with 2-trichloroacetylpyrrole at room temperature. The amide group is twisted by 7.4 (1)degrees from the plane of the pyrrole ring. In the crystal, molecules are linked by intermolecular N-H...O hydrogen bonds into chains extending along the c axis." /> <meta name="prism.number" content="4" /> <meta name="prism.volume" content="67" /> <meta name="prism.publicationDate" content="2011-04-01" /> <meta name="prism.publicationName" content="Acta Crystallographica Section E: Structure Reports Online" /> <meta name="prism.issn" content="1600-5368" /> <meta name="prism.section" content="organic compounds" /> <meta name="prism.startingPage" content="752" /> <meta name="prism.rightsAgent" content="med@iucr.org" /> <meta name="prism.endingPage" content="752" /> <meta name="prism.eissn" content="1600-5368" /> <meta name="keywords" lang="en" content="" /> <meta name="ROBOTS" content="NOARCHIVE,NOINDEX" />
IUCr bibJSON:
[
{
"type" : "metadata",
"namespaces" : {
"dc" : "http://purl.org/dc/elements/1.1/ ",
"prism" : "http://prismstandard.org/namespaces/1.2/basic/ ",
"bibo" : “http://purl.org/ontology/bibo/ "
}
},
{
"url" : "http://scripts.iucr.org/cgi-bin/paper?cv5056 ",
"author" : [
"Zheng, L.",
"Hu, F.",
"Zeng, X.C.",
"Li, K.P."
],
"abstract" : "The title compound, C11H14N2O5....",
"journal" : "Acta Crystallographica Section E: Structure Reports Online"
"bibo:issn" : "1600-5368",
"bibo:doi" : "10.1107/S1600536811007148",
"dc:rights" : "",
"dc:date" : "2011-04-01",
"dc:publisher" : "International Union of Crystallography",
"dc:language" : "en",
"dc:description" : "The title compound, C11H14N2O5...",
"dc:title" : "rac-Dimethyl 2-(1H-pyrrole-2-carboxamido)butanedioate",
"prism:number" : "4",
"prism:volume" : "67",
"prism:section" : "organic compounds"
"prism:startingPage" : "752",
"prism:endingPage" : "752",
"prism:publicationDate" : "2011-04-01",
"prism:eissn" : "1600-5368",
"prism:rightsAgent" : "med@iucr.org",
}
]
Atmospheric Chemistry and Physics BibTex:
@Article{acp-11-4679-2011,
AUTHOR = {Murphy, D. M. and Chow, J. C. and Leibensperger, E. M. and Malm, W. C. and Pitchford, M. and Schichtel, B. A. and Watson, J. G. and White, W. H.},
TITLE = {Decreases in elemental carbon and fine particle mass in the United States},
JOURNAL = {Atmospheric Chemistry and Physics},
VOLUME = {11},
YEAR = {2011},
NUMBER = {10},
PAGES = {4679--4686},
URL = {http://www.atmos-chem-phys.net/11/4679/2011/ },
DOI = {10.5194/acp-11-4679-2011}
}
Atmospheric Chemistry and Physics BibJSON:
[
{
"type" : "article",
"author" : [
"Murphy, D. M.",
"Chow, J. C",
"Leibensperger, E. M.",
"Malm, W. C.",
"Pitchford, M.",
"Schichtel, B. A.",
"Watson, J. G.",
"White, W. H."
],
"title" : "Decreases in elemental carbon and fine particle mass in the United States",
"journal" : "Atmospheric Chemistry and Physics",
"volume" : "11",
"year": "2011",
"number" : "10",
"pages" : "4679--4686",
"url" : "http://www.atmos-chem-phys.net/11/4679/2011/ ",
"doi" : "10.5194/acp-11-4679-2011",
}
]
J.ChemInf bibJSON (based on JChemInf RDF):
[
{
"type" :"metadata",
"namespaces" : {
"dc" : "http://purl.org/dc/elements/1.1/ ",
"dcterms" : "http://purl.org/dc/terms/",
"prism" : "http://prismstandard.org/namespaces/1.2/basic/ "
},
},
{
"url" : "http://www.jcheminf.com/content/3/1/17 ",
"bibjson:fulltext" : "http://www.jcheminf.com/content/pdf/1758-2946-3-17.pdf ",
"abstract" : "http://www.jcheminf.com/content/3/1/17/abstract/ ",
"title" : "ChemicalTagger: A tool for semantic text-mining in chemistry",
"author" : [
"Lezan Hawizy",
"David Jessop",
"Nico Adams",
"Peter Murray-Rust"
],
"journal" : "Journal of Cheminformatics 2011 3:17",
"dc:date" : "2011-5-16",
"dc:identifier" : "",
"dc:publisher" : "Chemistry Central Ltd",
"dc:rights" : "",
"dc:language" : "en",
"dc:format" : "text/html"
"prism:publicationName" : "Journal of Cheminformatics"
"prism:issn" : "1758-2946",
"prism:publicationDate" : "2011-5-16",
"prism:volume" : "3",
"prism:number" : "1",
"prism:startingPage" : "17",
"prism:copyright" : "2011 Hawizy et al;",
"prism:rightsAgent" : "reprints@biomedcentral.com",
}
]
## namespace prefixes used: @prefix rdf: . @prefix rdfs: . @prefix owl: . @prefix dc: . @prefix dcat: . @prefix void: . @prefix bibo: . @prefix cito: . @prefix foaf: . @prefix skos: . @prefix opmv: . @prefix time: . @prefix xsd: . ## metadata about the medline dataset the current record is in: a void:Dataset, dcat:Dataset ; ## licens terms of this (RDF) dataset dc:license ; ## the RDF generation finished at this time dc:modified "2011-05-08T15:23:45Z"^^xsd:dateTime ; ## it came from this medline XML file dc:source "medline11n0421" ; ## which can be obtained here (in theory, not yet) void:dataDump ; ## another way of expressing where it can be obtained dcat:distribution [ dc:description "bzip2 compressed N-Quads" ; a dcat:Distribution ; dcat:accessURL ] . ## information about the medline record in question: a foaf:Document; ## link back to the dataset dc:isPartOf ; ## license terms of this metadata dc:license ; ## various timestamps relating to dc:created "2000-06-28"^^xsd:dateTime ; dc:issued "2000-06-28"^^xsd:dateTime ; dc:modified "2004-11-17"^^xsd:dateTime ; ## provenance information dc:source "MEDLINE" ; opmv:wasGeneratedBy [ a opmv:Process ## we used version 1.3 of the medline software, and the ## indicated source dataset opmv:used , [ rdfs:label "medline11n0421" ] ; ## it was me that did this conversion opmv:wasControlledBy ; ## at this time opmv:wasPerformedAt [ a time:Instant ; time:inXSDDateTime "2011-05-08T15:23:45Z"^^xsd: dateTime ] ; ] ; ## this metadata record is about this article foaf:primaryTopic . ## some further information about me, the person who did the conversion a foaf:Agent ; foaf:mbox ; foaf:name "William Waites" . ## some further information about the software that was used dc:version "1.3" ; rdfs:label "Go Medline 1.3" . ## this is the journal that the article was published in a bibo:Journal ; dc:identifier "0077-8923", "7506858" ; dc:title "Annals of the New York Academy of Sciences" ; bibo:issn "0077-8923" ; ## we can use the ISO abbreviation to build up a concept ## of standard names for journals skos:prefLabel "Ann. N. Y. Acad. Sci." . ## information about the actual article a bibo:AcademicArticle ; dc:title "Paradoxical phosphorylation of skeletal muscle glycogen synthase by in vivo insulin in very lean young adult rhesus monkeys." ; dc:published "1999-18" ; dc:language "eng" ; ## this article was published in this issue... dc:isPartOf [ a bibo:Issue ; bibo:volume "892" ; dc:published "1999-18" ; dc:spatial "UNITED STATES" ; ## .. which is part of this journal dc:isPartOf ; ] ; ## .. on these pages dc:extent "247-60" ; ## and here are the authors dc:creator [ a foaf:Person ; foaf:familyName "Ortmeyer" ; foaf:givenName "H K" ; foaf:name "Ortmeyer, H K" ], [ a foaf:Person ; foaf:familyName "Bodkin" ; foaf:givenName "N L" ; foaf:name "Bodkin, N L" ], [ a foaf:Person ; foaf:familyName "Hansen" ; foaf:givenName "B C" ; foaf:name "Hansen, B C" ] .